Research
Magnetic tweezers
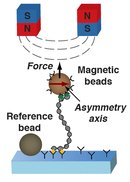 Magnetic tweezers are a powerful technique to apply both forces and torques to individual macromolecules. We use magnetic tweezers to study DNA, RNA and proteins. Magnetic tweezers measurements provide fundamental information about the mechanical properties of macromolecules and can reveal conformational transitions under external forces and torques. They are also a great tool to probe interactions between macromolecules as well as with small-molecule drugs.
Magnetic tweezers are a powerful technique to apply both forces and torques to individual macromolecules. We use magnetic tweezers to study DNA, RNA and proteins. Magnetic tweezers measurements provide fundamental information about the mechanical properties of macromolecules and can reveal conformational transitions under external forces and torques. They are also a great tool to probe interactions between macromolecules as well as with small-molecule drugs.
We have developed novel tweezers that can directly measure torque (so-called magnetic torque tweezers, MTT) and directly observe changes in the twist of nucleic acid tethers (termed freely-orbiting magnetic tweezers or FOMT). These developments enhance the capabilities of conventional magnetic tweezers and provide orthogonal dimensions of single-molecule measurements and control.
Recently, we have pioneered strategies that enable ultra-stable and versatile attachment of proteins in magnetic tweezers and used these capabilities to investigate binding interfaces and conformational transitions of proteins under force.
Small-angle X-ray scattering
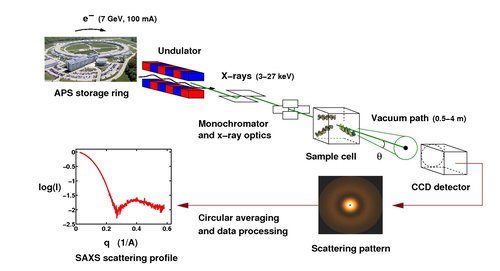
Small-angle X-ray scattering (SAXS) can probe the (low-resolution) structure, conformational changes, and interactions of macromolecules in solution, under almost arbitrary solution conditions, from physiological to highly denaturing. A schematic of a synchrotron SAXS measurement is shown below (the parameters and data shown are for the Advanced Photon Source at Argonne National Lab near Chicago). In addition to using synchrotron radiation, we employ an in-house X-ray source for SAXS measurements.
Atomic force microscopy and automated image analysis
Atomic force microscopy (AFM) enables quantitative imaging of biological macromolecules with (sub-)nanometer resolution. By imaging DNA, nucleosomes (the first compaction step of DNA) and other protein complexes, we want to get new insights into their structural properties and biological functions. To obtain statistically relevant and reliable data, we additionally develop a high-throughput automated AFM image analysis toolbox that combines both classical and machine learning analysis approaches.


